About the OBF
The Open Bioinformatics Foundation (OBF) is a non-profit, volunteer-run group that promotes open source software development and Open Science within the biological research community. Membership in the OBF is open to anyone who wants to help promote open source or open science in a biological field.
OBF runs the annual Bioinformatics Open Source Conference (BOSC). BOSC 2023 took place July 24-25, 2023, as part of ISMB/ECCB 2023 in Lyon, France and online. It was followed by CoFest 2023. BOSC 2024 will be July 15-16, 2024, at ISMB 2024 in Montréal, Canada.
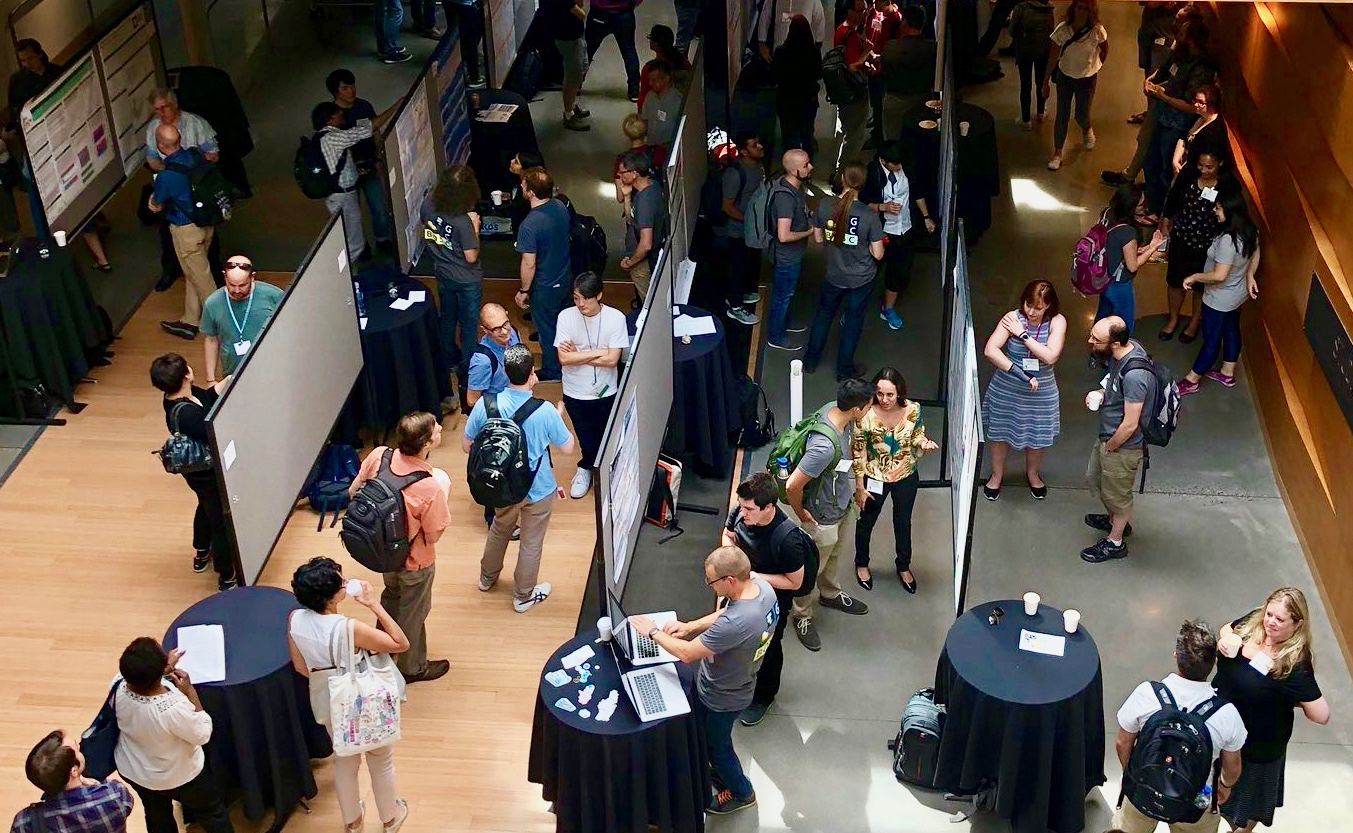


OBF Event Awards
The OBF Event Fellowship program aims to increase diverse participation at events promoting open source bioinformatics software development and open science in the biological research community.
- What are the selection criteria?
- What does the fellowship cover?
- What do you require of applicants?
- Who will review the applicants?
Board of directors
-
 At-large Member
At-large Member -
 At-large Member
At-large Member -
 At-large member
At-large member -
 At-large Member
At-large Member -
 President
President -
 At-large Member
At-large Member -
 Secretary
Secretary -
 Treasurer
Treasurer -
 At-large Member
At-large Member